 Īs attractive as it might seem financially to run just one microarray for each “class” of samples (of the same phenotype, time-point, or tissue type) under consideration, replicates are essential for providing meaningful results. However, clever design can somewhat reduce the required number of arrays. Dye swapping imposes additional costs in both the number of arrays and the types of data analyses possible. “Dye-swap” experiments, in which the same pairs of samples are compared twice with the labeling colors swapped, can permit the computational removal of such bias. Challenges include ensuring that all samples can be compared to the appropriate controls and avoiding any biases introduced by the different labeling. Affymetrix arrays are inherently single-channel, though some associated analysis tools facilitate pair-wise comparisons.ĭesign issues for two-color arrays are more complex. Agilent and NimbleGen arrays can be run using either one or two channels. cDNA arrays typically involve two channels. A recent comparison of single- and two-color methods on the same platforms found good overall agreement in the data produced by the two methods. Single-color arrays allow for more flexibility in analysis, while two-color arrays can control for some technical issues by allowing a direct comparison in a single hybridization. There are tradeoffs between the two approaches. Slightly different oligonucleotide array platforms are manufactured by companies such as Affymetrix, Agilent, and NimbleGen (see Text S1 and Table S1 for further discussion).Ī major design question is whether to measure the expression levels from each sample on a different microarray (using single-color, or single-channel, arrays), or instead to compare relative expression levels between a pair of samples on each microarray (two-color or two-channel arrays). 
While the former may be less expensive because they can be manufactured in the lab or at institutional core facilities, the latter may outperform the former in terms of number of spots per array and the spots' homogeneity. The main distinction is whether essentially full-length transcripts are printed onto slides (cDNA microarrays) or the desired-typically shorter-oligonucleotides are synthesized in situ (oligonucleotide arrays). 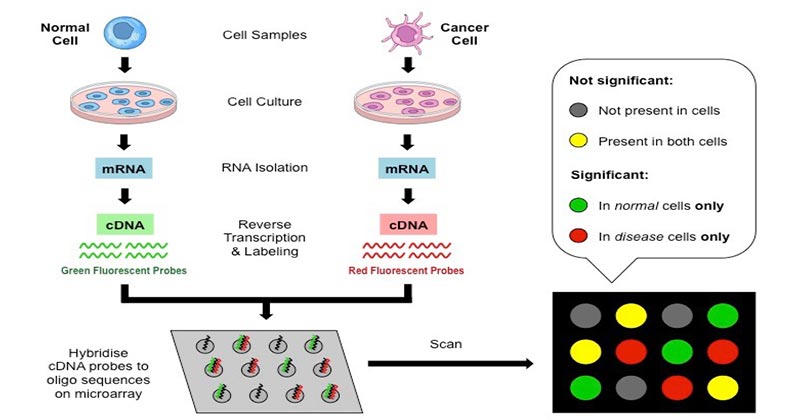
Design issues depend in part on the exact array technology used, and indeed, choosing an array technology is often the first design choice. Careful experimental design is crucial for a successful microarray experiment, , yet this important step is often shortchanged. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
